How Complement Fight Infections?
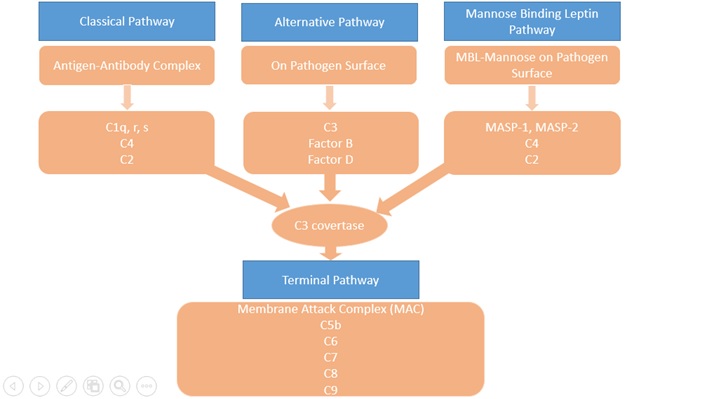
The complement system consists of more than 30 proteins, present in blood and tissues, as well as other proteins anchored on the surfaces of cells. The primary functions of the complement system are to protect against infection, to remove particulate substances, (like damaged or dying cells, microbes, or immune complexes), and to help modulate adaptive immune responses. As part of the innate immune system, the complement acts immediately to start the process of removal and resolution of the problem. The complement works with the inflammatory cells of the innate immune system and those of adaptive or acquired immunity. Complement proteins in the circulation are not activated until triggered by an encounter with a bacterial cell, a virus, an immune complex, damaged tissue, or other substance not usually present in the body. There are three major activations of complement pathways; classical (CP), alternative(AP), and leptin pathway(LP) summarised in Figure 1. The Classical Pathway (CP) is activated primarily by immunoglobulins that are bound to antigens. Aggregates of immunoglobulins such as cryoglobulins also activate the CP. Components of the CP are C1q, C1r, C1s, C2, and C4. The Leptin Pathway (LP) is initiated by Mannose-binding Leptin (MBL) or one of the Ficolins to a target molecule. Both MBL and ficolins are present in the complex with MBL-associated serine protease(MASP) and activate C4 and C2. The Alternative Pathway (AP) is initiated by fragments of the complement component C3. Factor B, Factor D, and properdin also involved in the AP. Properdin, the only positive regulator in the complement system available in AP. Makes it possible for the amplification loop of the alternative pathway to set up a very efficient mechanism for putting lots of C3b onto the surface of the activating cells, protein complexes, or particles in the immediate vicinity of the activation site. These three pathways converge at the component C3. Although each branch is triggered differently, the common goal is to deposit clusters of C3b on a target. This deposition provides for the assembly of the membrane attack complex (MAC).
The Terminal Pathway (TP) is the final set of steps in the complement activation process that forms a membrane lesion or hole (membrane attack complex or MAC) that kills susceptible bacteria or other cells that activate the complement on their surfaces. The TP is dependent upon at least one of the other pathways to initiate the process, that it then completes. The components of the TP are C3, C5, C6, C7, C8, and C9. A form of the MAC, called the Terminal Complement Complex (TCC) can be found in the circulation after complement activation occurs and makes a useful laboratory marker for complement activation.
Figure 1: Complement Pathways
Introduction
The complement system is part of the immune system, whose major function is to protect the host from infections. As a part of the defense, both the innate and the adaptive immune systems are operative through several mechanisms. The complement system comprises an important part of the innate defense and some by bridging the innate and adaptive immunity (Dunkelberger et al 2010). It promotes the inflammatory response, eliminates pathogens, and enhances the immune response. Deficiencies in the complement cascade can lead to overwhelming infections and sepsis (JE Figueroa, Denssen 1991). In addition to playing an important role in host defense against infection, the complement system is a mediator in both the pathogenesis and prevention of immune complex diseases, such as systemic lupus erythematosus.
The importance of complement in defense against bacterial infections has become clear with some role against fungal and parasitic infection (Hirsch RL et al 1981). The ability of Mannose-binding lectin to bind to HIV has been shown in an in-vitro study (Saifuddin et al 2000).
Complement deficiencies are rare. However, they are poorly characterised clinically and have been difficult to detect, so are likely to be significantly underdiagnosed. Complement deficiencies form about 1-10% of all primary immunodeficiency disorders (Grumach et al 2014). Complement deficiencies can be inherited or acquired. The genetic deficiency of components involved in the classical pathway (C1q, C1r/s, C2, C4) tend to be linked with autoimmune diseases(Bryan et al 2014), whereas C5 to C9 may have enhanced susceptibility to meningococcal disease.
Complement Deficiencies and Associated Infections
There is a wide variation of infections associated with complement deficiency depending on which complement protein and activation pathway is affected(Skattum et al 2011). Some complement deficiencies carry a risk of infection, others are mainly associated with autoimmune diseases. Some genetic and molecular defects have been identified to be associated with complement deficiencies. The deficiencies in the alternative pathway and the terminal pathway are commonly associated with markedly increased susceptibility to Neisserial infections. Properdin deficiency (PD) has been reported in cases of severe meningococcal infections (Fijen et al 1999), recurrent otitis media, and pneumonia (Schejbel et al 2009). PD has been identified to have an X-linked inheritance. Around 500 published cases of PD with Neisserial infections worldwide. Neisserial meningitidis meningitis and Streptococcal pneumonia also have been reported in complete Factor D deficiency. The TP component deficiencies have been reported in association with Neisserial infections. Deficiency of C5, C6, C7 and C8 has less than 1000 reported cases of Neisserial meningitis and sepsis, while C9 deficiencies are more common in Japanese populations with prevalence around 1:1000 (Fukumori et al 1989)(Nagata et al 1989). In LP, MBL deficiencies have been linked to causing recurrent infections in children(Super et al, 1989). Around 10- 15% of Caucasians carry the MBL gene that shows a deficient concentration of functional MBL. Another rare deficiency MASP- deficiency, which reported in an adult with autoimmune symptoms and recurrent respiratory infections (Stengaard-Pedersen 2003) and Ficolin-3 deficiency has been described in patients with a recurrent respiratory infection and cerebral abscess (Munthe-Fog et al). Increased susceptibility to encapsulated bacteria, S. pneumonia, and N. meningitidis is a common feature of CP deficiencies. CP deficiency is also associated with autoimmune disorders, like systemic lupus erythematosus. Few identified genetically determined deficiencies are C2D, C1q, and C4 (Truedsson et al and Sjoholm et al). C2D is associated with abnormalities in serum immunoglobulin levels and associated with invasive encapsulated bacteria infection and autoimmune disease (Jonsson et al). Other uncommon deficiencies such as factor H deficiency, the factor I deficiency which is associated with susceptibility to encapsulated bacteria along with autoimmune diseases such as glomerulonephritis and vasculitis (Nilsson et al, Nita et al). C1 INH deficiency with a prevalence of approximately 1 in 50,000 presents as hereditary angioedema. Complement deficiencies and associated genetic disorders and clinical manifestations are summarized in Table 1.
Table 1 - Complement deficiencies and clinical associations.
| Proteins |
Genetics |
Frequency |
Infections |
Clinical features/manifestations |
Other associations |
| C1q |
Autosomal recessive |
50-100 |
Sepsis, meningitis, pneumonia,
Streptococcus (S) pneumoniae, Neisseria meningitidis |
SLE like symptoms (malar discoid rash, oral ulcers, arthritis (less common)[1]. |
SLE |
| C1r/C1s |
Autosomal recessive |
<20 |
Encapsulated bacterial pneumonia, meningitis |
SLE like symptoms, glomerulonephritis in 50% patients[2] |
SLE |
| C4 |
Autosomal recessive |
20 -50 |
Sepsis, meningitis, pneumonia,
Streptococcus (S) pneumoniae, Neisseria meningitidis |
Glomerulonephritis in 50% patients[2] |
SLE |
| C2 |
Autosomal recessive |
1/20 000 |
S.pneumonia, staphylococcus aureus, H. Influenza |
SLE like symptoms like arthritis, malar rash, discoid rash, photosensitivity and serositis[2] |
SLE |
| C3 |
Autosomal recessive |
20-50 |
Respiratory tract infection, meningitis |
Recurrent systemic infections, rheumatic and renal problems.[3] |
Immune-complex disease |
| Factor D |
Autosomal recessive |
<50 |
Meningitis, sepsis due to Neisseria |
Recurrent infections, especially Neisseria [4] |
|
| Properdin |
X-linked |
100-500, common in Caucasians[5] |
Meningitis, sepsis due to neisseria |
Neisseria infections[5] |
|
| MBL |
Common polymorphism in exon 1 |
5-15% of Caucasian |
Respiratory infections |
Respiratory infections, otitis media, chronic diarrhoea [6] |
cardiovascular |
| MASP-2 |
Autosomal recessive |
<20 |
Respiratory infections |
|
autoimmune |
| Ficolin 3(H-ficolin) |
Autosomal recessive |
1 |
Respiratory infections
Cerebral abscess, Streptococci. H influenza. Pseudomonas aeruginosa |
SLE like symptoms, recurrent infections [7] |
thrombocytopenia |
| C5 |
Autosomal recessive |
20-50 |
Meningitis due to neisseria |
|
| C6 |
Autosomal recessive |
50-100 |
Meningitis due to Neisseria |
Recurrent meningococcal infections [8] |
|
| C7 |
Autosomal recessive |
50-100 |
Meningitis due to Neisseria |
|
| C8 |
Autosomal recessive |
50-100 |
Meningitis due to Neisseria |
|
| C9 |
Autosomal recessive |
1/1000 in Japanese |
Meningitis due to Neisseria |
|
| Factor H |
Autosomal recessive |
<50 |
Recurrent pyogenic infections N.meningitidis, H influenzae |
|
Membrano proliferative glomerulonephritis, HUS |
| Factor I |
Autosomal recessive |
5% Caucasian |
Recurrent pyogenic infections N.meningitidis, H influenzae |
|
Immune complex related disease |
| C1-INH |
Autosomal dominant |
1/50000 |
|
Facial edema, edema of upper respiratory tract[9] |
HAE |
| CD59, CD55 |
Somatic mutations of PIG-A |
1-2/1,000,000 |
Cytopenias |
Pallor, fever, tiredness and haemoglobinuria[10] |
Thrombosis, hematopoietic cytopenia |
| CR3/CD4 (LAD 1) |
Autosomal recessive |
1/1,000,000 |
S. aureus, Gram-negative bacteria |
Recurrent infections, delayed separation of umbilical cord, dental problems like periodontitis [11] |
|
Test for Complement Deficiencies
For the evaluation of particular pathways of the complement system, functional tests have been developed. The ‘total hemolytic complement’ or CH50 is based on a hemolytic assay in which a series of patient serum dilutions are incubated with sheep erythrocytes covered with specific IgM, activating the classical complement pathway (Wen L et al). The reciprocal of the serum dilution which causes the lysis of 50% of the erythrocytes is calculated to quantify the amount of active complement. For example, if 50% of the sheep erythrocyte are lysed at a serum dilution of 1:200, this equal to a CH50 value of 200 units/ml. Since the formation of membrane attack complexes (MACs) is necessary for the lysis of the erythrocytes and requires all components of the classical and common terminal complement pathway (C1 through C9), a deficiency or defect in any of these components would result in a significantly lowered CH50. Several variants of this test are in use example one being based on liposomes containing enzymes as a read-out to simplify the quantification of lysis using a standard analyzer (Wen et al). Thus, reference values of CH50 may vary depending on the system that is used.
A similar test, the AH50 is used to assess the alternative complement pathway (properdin, factor B, D, and C3) in conjunction with the common terminal pathway (C5 through C9) (Wen et al). Serum dilutions are incubated with rabbit erythrocytes in a buffer that inhibits the classical or leptin pathway. If the alternative pathway is intact, a properdin-stabilized C3 convertase (C3bBb) can form on the surface of the erythrocytes and allow the assembly of C5 convertase and MACs, resulting in lysis which can be quantified.
Using both CH50 and AH50 can help to map the complement deficiency to a specific pathway. If the classical pathway is affected, the CH50 value will be low and the AH50 value will be normal. The opposite is true for a defect in the alternative pathway, and if both CH50 and AH50 are low, the abnormal component is likely to be part of the common terminal pathway. However, CH50 and AH50 can also be low due to errors in sample handling because several complement components are unstable (Lock RJ et al). In case of such a result, the quality of the specimens should be controlled and the test might need to be repeated, if in doubt.
To test the leptin pathway, a mannan-based ELISA can be used. MBL from the patient’s serum binds the mannan-coated wells, leading to MASP-activation and cleavage of C4. The fragments C4b and C4d can then be quantified using enzymes-linked antibodies (Wen et al).
In order to eventually identify the deficiency, serum levels of individual complement factors can be measured by immunochemical assays based on immunoprecipitation and nephelometry (measurement of turbidity caused by antigen-antibody complexes). Such tests are readily available for C3 and C4. However, quantification of other complement components might require the help of specialized laboratories. One major drawback of this method is the dependence on the quality of antibodies and standards used. Moreover, functional defects i.e non- functional complement factors with normal serum levels are not detected. Genetic testing, by sequencing exons coding for complement components, avoids these problems.
Management
Causative therapies are currently unavailable and often unnecessary as standard treatments of complement deficiencies. Replacement of the respective complement components, either recombinant or plasma products, would have too many disadvantages: most complement factors have a short half-life and would require frequent intravenous administration which would be disproportionately inconvenient for the patients. Moreover, they would be costly, possibly immunogenic and plasma products would come with a theoretical risk of transmitting infectious agents. Even in the case of acute infection, replacement therapies are redundant since the patients usually respond to treatment with antibiotics.
Resistance to infections in complement deficient patients can be increased most efficiently by active immunisation. In particular, vaccination against encapsulated bacteria is recommended (Jonsson G et al). These include Haemophilus influenza type b, Streptococcus pneumonia, and the main Neisseria meningitidis serotypes A, C, W135, Y, and possibly B, with the introduction of the new serotype B vaccine. Subsequently, the antibody titers should be controlled to ensure that they are protective. More severe cases with recurrent or persistent serious infections might require prophylactic antibiotic therapy (Kuruvilla M et al)
Concluding Remarks
The role of complements in the immune system has expanded dramatically. It is a well-characterized and an evolving component of host defense, impairment of which leads to susceptibility to infection. Complement represents a cornerstone of the innate defense against infection and provides a vital first-line barrier by activation of proteolytic cascades which leads to the identification and persecution of the surface antigens, identified as foreign, and allows the complement to contain, control, and finally clear invading microorganisms. On top of these important contributions to innate immunity, complement plays a vital role in shaping adaptive immune responses, functionally integrating it into the ability of the host to combat invasion from a wide range of pathogens. The wide range of clinical manifestations of complement deficiencies reflects the complexity of the complement system. As more individuals are identified with specific complement deficiencies, our understanding of how the complement system functions in innate and adaptive immunity will evolve. In conclusion, complement is a multifaceted and robust effector, which bridges the innate and adaptive immune systems. It is vital to the host defense, and the extent of its influence is becoming increasingly appreciated as additional information regarding the far-reaching effects of its activation is uncovered. Further studies should produce significant knowledge and contribute to our understanding of host defense as an integrated process and the roles complement plays in bridging innate and adaptive immunity.
1. Stegert M, Bock M, Trendelenburg M. Clinical presentation of human C1q deficiency: How much of a lupus? Mol Immunol. 2015 Sep;67(1):3-11. doi: 10.1016/j.molimm.2015.03.007. Epub 2015 Apr 3. PMID: 25846716.
2. Macedo AC, Isaac L. Systemic Lupus Erythematosus and Deficiencies of Early Components of the Complement Classical Pathway. Front Immunol. 2016 Feb 24;7:55. doi: 10.3389/fimmu.2016.00055. PMID: 26941740; PMCID: PMC4764694.
3. https://rarediseases.org/mondo-disease/complement-component-3-deficiency/
4. Biesma DH, Hannema AJ, van Velzen-Blad H, Mulder L, van Zwieten R, Kluijt I, Roos D. A family with complement factor D deficiency. J Clin Invest. 2001 Jul;108(2):233-40. doi: 10.1172/JCI12023. PMID: 11457876; PMCID: PMC203023.
5. Linton SM, Morgan BP. Properdin deficiency and meningococcal disease--identifying those most at risk. Clin Exp Immunol. 1999 Nov;118(2):189-91. doi: 10.1046/j.1365-2249.1999.01057.x. PMID: 10540177; PMCID: PMC1905414.
6. Turner MW. Deficiency of mannan binding protein--a new complement deficiency syndrome. Clin Exp Immunol. 1991 Oct;86 Suppl 1(Suppl 1):53-6. doi: 10.1111/j.1365-2249.1991.tb06208.x. PMID: 1934604; PMCID: PMC1554043.
7. Troldborg A, Steffensen R, Trendelenburg M, Hauser T, Winther KG, Hansen AG, Stengaard-Pedersen K, Voss A, Thiel S. Ficolin-3 Deficiency Is Associated with Disease and an Increased Risk of Systemic Lupus Erythematosus. J Clin Immunol. 2019 May;39(4):421-429. doi: 10.1007/s10875-019-00627-2. Epub 2019 May 1. PMID: 31044336.
8. Fischer E. Hereditäre Komplmentdefekte [Hereditary complement deficiencies]. Hautarzt. 1982 Feb;33(2):65-72. German. PMID: 6281215.
9. Sobotkova M, Zachova R, Hakl R, Kuklinek P, Kralickova P, Krcmova I, Hanzlikova J, Vachova M, Bartunkova J. Acquired Angioedema with C1 Inhibitor Deficiency: Occurrence, Clinical Features, and Management: A Nationwide Retrospective Study in the Czech Republic Patients. Int Arch Allergy Immunol. 2021;182(7):642-649. doi: 10.1159/000512933. Epub 2021 Jan 20. PMID: 33472202; PMCID: PMC8315685.
10. Khan M, Ahmad SQ, Bashir M, Ahmed P, Ayyub M. Clinico-Haematological Features of Paroxysmal Nocturnal Haemoglobinuria. J Coll Physicians Surg Pak. 2017 Jan;27(1):53-54. PMID: 28292372.
11. Justiz Vaillant AA, Ahmad F. Leukocyte Adhesion Deficiency. 2023 Jul 3. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30969592.